SNP data mining heteroplasmy mitochondria. In conclusion, our study provides important guidelines for future studies that intend to use either exome sequencing or RNAseq data to infer mitochondrial SNPs and heteroplasmy. Furthermore, we examined three alignment strategies and found that aligning reads directly to the mitochondrial reference genome or aligning reads to the nuclear and mitochondrial references genomes simultaneously produced the best results, and that aligning to the nuclear genome first and afterwards to the mitochondrial genome performed poorly. If using the same reference, the index step only needs to be done once. However, the lower false discovery rate makes exome sequencing a better choice for heteroplasmy detection than RNAseq. Aligning reads using STAR is a two step process: Create a genome index Map reads to the genome Step 1. It uses sequential maximum mappable seed. Process all samples: Align all reads from two conditions (WT and SNF2) In the previous steps, we learned how to do quality control and read alignment using one WT fastq file as an example. 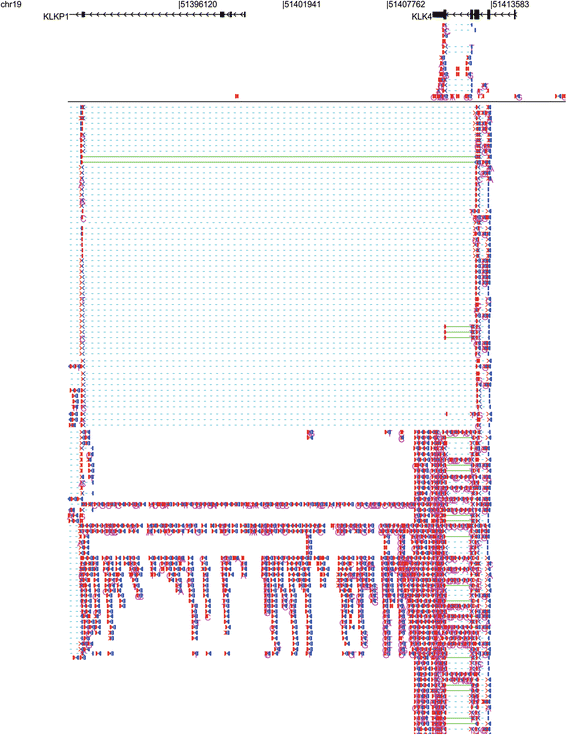
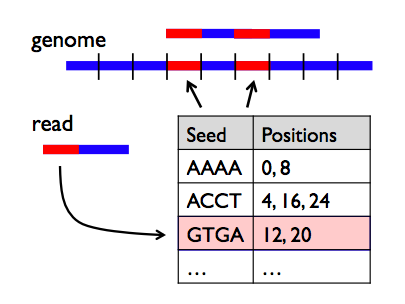
STAR is an ultrafast universal RNA-seq aligner. We see in the BAM track that reads are spiced across the introns and that coverage track that read coverage in the area of the intron is missing as expected. The indexing scheme is called a Hierarchical Graph FM index (HGFM). We found that exome and RNA sequencing can accurately detect mitochondrial SNPs. It is a fast and sensitive alignment program for mapping next-generation sequencing reads (both DNA and RNA) to a population of human genomes (as well as to a single reference genome). Mitochondrial targeted sequencing was used as the gold standard to compute the validation and false discovery rates of SNP and heteroplasmy detection in exome and RNAseq data. Five breast cancer cell lines were sequenced through mitochondrial targeted, exome, and RNA sequencing. We designed a study to evaluate the practicability of mtDNA variant detection using exome and RNA sequencing data. Recently, RNAseq data have also been suggested as a good source for alternative data mining, but whether mitochondrial variants can be detected from RNAseq data has not been validated. MacVector has a software plugin called Assembler that integrates directly into the DNA sequence analysis toolkit and provides DNA sequence assembly functionality. However, the effectiveness and practicality of this approach have not been tested. In addition to performing specific mitochondrial targeted sequencing, an increasingly popular alternative approach is using the off-target reads from exome sequencing to infer mtDNA variants, including single nucleotide polymorphisms (SNPs) and heteroplasmy. The rapid progress in high-throughput sequencing has significantly enriched our capacity for studying the mitochondrial DNA (mtDNA).
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
